- Norton et al. Am J Obstet Gynecol. 2012 Aug;207(2):137.e1-8.
- Nicolaides et al. Am J Obstet Gynecol. 2012 Nov;207(5):374.e1-6.
- Ashoor et al. Ultrasound Obstet Gynecol. 2013 Jan;41(1):21-5.
- Verweij et al. Prenat Diagn. 2013 Oct;33(10):996-1001.
- Ashoor et al. Am J Obstet Gynecol. 2012 Apr;206(4):322.e1-5.
- Sparks et al. Am J Obstet Gynecol. 2012 Apr;206(4):319.e1-9.
- Gil et al. Fetal Diagn Ther. 2014;35:204-11.
- Nicolaides et al. Fetal Diagn Ther. 2014;35(1):1-6.
- Hooks et al. Prenat Diagn. 2014 May;34(5):496-9.
- Sparks et al. Prenat Diagn. 2012 Jan;32(1):3-9.
- Norton M, et al, N Engl J Med. 2015 Apr 23;372(17):1589-97.
- Data on file, Ariosa
- Stokowski, et al. N Engl J Med 2015 Apr 23;37217:1589-97
- White K, et al. Performance of targeted cfDNA analysis with microarray quantitation for assessment of fetal sex and sex chromosome aneuploidy risk. Presented at ACMG, Tampa FL March 2016
- ACOG Practice Bulletin No. 77. Obstet Gynecol 2007;109:217-27.
- http://phprimer.afmc.ca/Part2-MethodsStudyingHealth/Chapter6MethodsMeasuringHealth/Interpretingtestsonindividuals
- Wang E, et al. Prenat Diagn. 2013 Jul;33(7):662-6.
Extensive blinded, published validation data
The Harmony Prenatal Test is broadly studied, including 11 blinded published validation trials.1-11
In blinded prospective published trials including over 22 000 pregnant women from ages 18 to 50, Harmony demonstrated exceptional sensitivity and specificity.1-2,4,7,11
Harmony Microarray-based performance

Harmony is Validated for Pregnant Women of Any Age or Risk Categories and Trusted by Clinicians Worldwide
- Studied extensively in blinded prospective published trials including >22 000 pregnant women.1-2,4,7,11
- Clinicians in more than 100 countries have trusted Harmony to screen more than 3 000 000 pregnancies.12
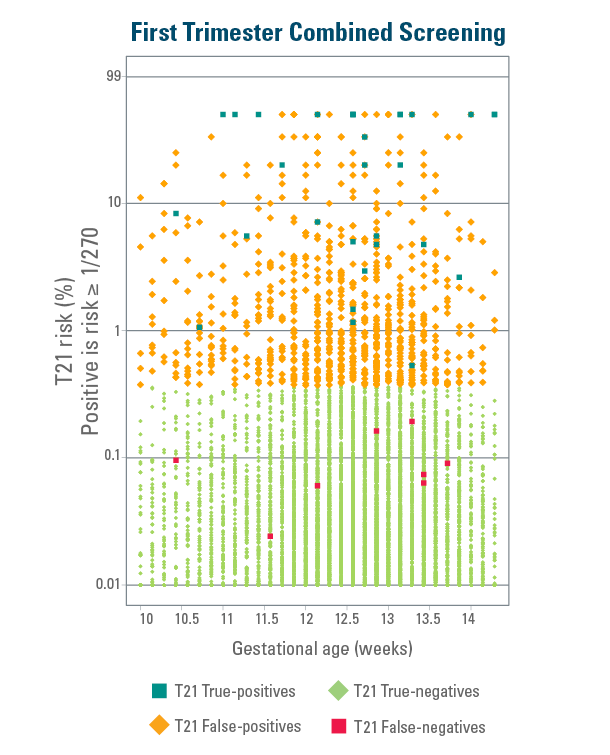
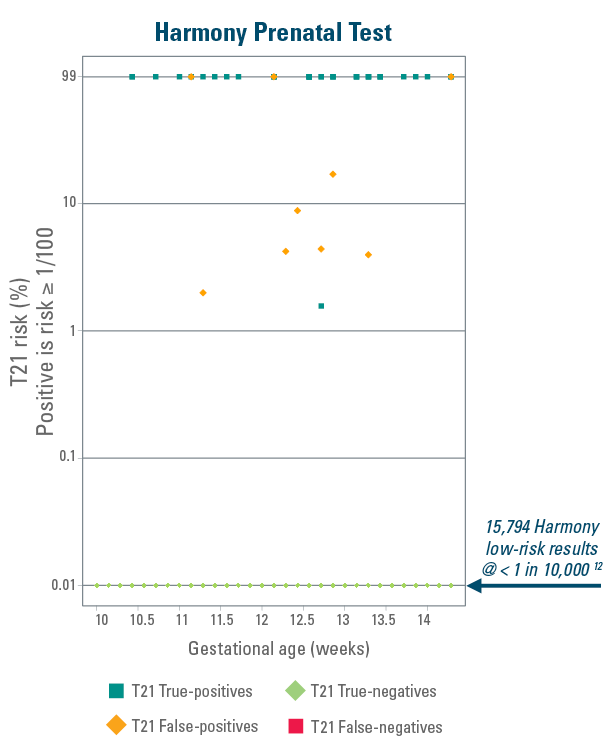
The Harmony Prenatal Test Shows Clear Difference Between High-Probability and Low-Probability Results12
The below graphs represent data from the blinded prospective head to head comparison study of First Trimester Screening (FTS*) against Harmony NIPT.11-12
Data represent 15 841 patients in a general pregnancy population.12
Harmony provides clear results for trisomy 21 risk, generating a wide separation between high-probability and low-probability values, with only extremely rare (less than 1 in 1000) false-positive results.12
*Serum PAPP-A, total or free ß-hCG & Nuchal Translucency
Exceptional positive predictive value (PPV)
Harmony has an extremely low false positive rate; less than 0.1% of all test takers get a false test answer. This is because the vast majority get a negative test result (low probability of trisomy).
Positive Predictive Value (PPV), on the other hand, indicates the probability that a positive test result is a correct one. PPV varies depending on the probability/prevalence of the event. Since trisomy 13, 18 and 21 have a higher prevalence among older pregnant women, the PPV also depends on the woman’s age in these cases.
As an example, the PPV for trisomy 21 is 91% in pregnancies in women who are 35 years of age, where the incidence of trisomy 21 is 1 in 250.1 In contrast, the PPV for KUB in the same population is 6%.15 Because the PPV for NIPT is never 100% , invasive sampling (amniocentesis or placental sampling) is always required to know for sure.
Below is a table with examples of how the PPV differs between the different chromosomal abnormalities, and the woman’s age, for Harmony NIPT:
| age 30 | age 33 | age 35 | age 40 | |
| T21 | 78,4% | 86,2% | 91,2% | 98,0% |
| T18 | 70,7% | 76,9% | 83,1% | 96,0% |
| T13 | 46,1% | 54,8% | 64,2% | 86,5% |
| T21/T18/T13 (combined) | 72,5% | 80,9% | 87,3% | 96,9% |
| Sex chromosome triploidies | 36,4% | 36,4% | 36,4% | 36,4% |
| Monosomy X | 28,6% | 28,6% | 28,6% | 28,6% |
| 22q11.2 | 11,0% | 11,0% | 11,0% | 11,0% |
Based on prevalence of 22q11.2 = 1/1524*
* Dar P, et al. Multicenter prospective study of SNP-based cfDNA for 22q11.2 deletion in 18,289 pregnancies with genetic confirmation [abstract]. Society for Maternal-Fetal Medicine. Virtual Meeting. Oral Presentation 67. Jan. 25-30, 2021.
Positive predictive value (PPV) and incidence
PPV depends on the incidence of the condition being tested. The rarer the condition, the higher the probability a positive result is a “false positive”.16
Testing for extremely rare conditions (1 in 10 000 incidence rate) in a routine genetic test may:
- Increase the cumulative false-positive rate of the entire test
- Substantially lower the overall PPV of the test
The percentage of fetal cfDNA is measured in every sample, and reported. If the fetal fraction is too low for reliable analysis, below 4%, no results will be reported but a redraw will be offered. The vast majority of samples greater than 10 weeks gestation contain an adequate fetal cfDNA proportion to allow for reliable clinical results. Wang et al showed that less than 2% of pregnant women had insufficient fetal cfDNA amounts for testing on the first blood draw17. The majority of the women had more than 4% fetal fraction of cfDNA on the second blood draw. Increasing maternal weight is associated with lower fetal fraction of cfDNA.



